The following protocol describes combined surface and intracellular protein staining with TotalSeq™-B antibodies, with or without TotalSeq™-B hashtag antibodies using the Single Cell 3’ v3.1 Feature Barcoding technology from 10x Genomics. This protocol has been optimized using liquid, single-antibody TotalSeq™ conjugates. We have not validated the use of TotalSeq™ Universal cocktails or TBNK cocktails at this time.
Disclaimer:
This protocol has been validated solely for the detection of surface or intracellular protein expression with the corresponding TotalSeq™-B antibodies. 10x Genomics has not validated this protocol and will not provide support to customers using this protocol. Gene expression data quality is likely to be negatively impacted when using this protocol. BioLegend does not guarantee high quality gene expression data with its use, and we are not able to provide sample datasets. Thus, users are advised to fully validate their application if they desire to obtain high quality data that includes surface proteins, intracellular proteins, and RNA expression data.
Only BioLegend clones validated using the reagents listed below are guaranteed to work as intended with this protocol. Staining of intracellular molecules can sometimes require specific fixation and permeabilization conditions that are not conducive between all clones. Clones not validated using this protocol should first be tested using this protocol to ensure staining is valid.
Reagent and Instrument List
- Fixation Buffer (BioLegend, Cat. No. 420801)
- Intracellular Staining Permeabilization Wash Buffer (10x) (BioLegend, Cat. No. 421002)
- Human TruStain FcX™ (Fc Receptor Blocking Solution) (BioLegend, Cat. No. 422301)
- TruStain FcX™ PLUS (anti-mouse CD16/32) (BioLegend, Cat. No. 156603)
- True-Stain Monocyte Blocker™ (BioLegend, Cat. No. 426101)
- Cell Staining Buffer (BioLegend, Cat. No. 420201)
- Buffer EB (Qiagen, Cat. No. 19086)
- Flowmi™ Cell Strainer (Bel-Art, H-B Instrument, Cat. No. H13680-0040)
- 12 x 75 mm Falcon™ Round-Bottom Polystyrene Tubes (Fisher Scientific, Cat. No. 14-959-1A or equivalent)
- Deionized water (DI water)
- Low Protein Binding Microcentrifuge Tubes (ThermoFisher Scientific, Cat. No. 90410 or equivalent)
Best Practices
Cell Washing
- When washing cells, it is extremely important that care is taken to thoroughly decant the wash buffer, and, that upon the addition of new wash buffer that the cell pellet is resuspended either with pipette mixing or gentle vortexing. When decanting, pour off the wash buffer in a single, confident and firm, but, not overly forceful motion. After decanting, continue holding the tube inverted. Any droplets remaining on the lip of the tube can be gently dabbed on a clean paper towel before returning the tubes to an upright position. This technique should be used during all cell washes.
Antibody Staining/Working Concentration
- Our liquid, single-antibody TotalSeq™ conjugates require optimization of staining concentration (titration) to obtain best results, as performed by the end user of the antibodies. This includes TotalSeq™ antibodies targeting intracellular proteins of interest.
- Antibody titration for sequencing-based applications using TotalSeq™ antibodies is best performed using matching procedures as much as possible. This means titration-by-sequencing if possible. In the absence of titration-by-sequencing, it may be possible to replicate this protocol using fluorescent antibodies of the same clone and titrate via flow cytometry. Optimization of antibody staining concentration is essential to obtain good quality data in antibody-based applications such as surface and intracellular staining using TotalSeq™ antibodies.
Protocol
Cell Surface Staining
- Prepare cell suspension with preferred or recommended method
Notes:
- This protocol has been optimized using fresh human PBMCs isolated using density gradient centrifugation. Whole blood or lysed whole blood is not recommended. If using cells isolated with a different procedure, users may need to verify the antibody staining pattern using alternative methods.
- BioLegend has not tested this protocol using single-cell suspensions derived from enzymatically digested tissue. Enzymatic digestion may result in alterations of surface protein epitopes and impact staining with TotalSeq™ antibodies. Optimization of staining conditions and concentrations may be required.
- Depending on your experimental set up, you may find abundant intracellular targets such as cytokines. However, it is always advisable to include a biological positive control. Activated cells can be prepared from in vivo-stimulated tissues or from in vitro-stimulated cultures (e.g. mitogen or LPS stimulation). For cytokine and chemokine detection, it is critical to include a protein transport inhibitor such as Brefeldin A (BioLegend Cat. No. 420601) or Monensin (BioLegend Cat. No. 420701) in the last 4-6 hours of cell culture activation. For details on stimulation methods, please see our stimulation guide for cytokines/chemokines.
- Count and assess cell viability
2.1 Carefully count all cells to ensure accurate quantitation and assess cell viability. Ideal cell viability is ≥ 95%.
2.2 Low cell viability is associated with generation of poor data and is not ideal for single-cell sequencing experimentation. If low cell viability is observed, users may need to enrich live cells or repeat cell suspension preparation.
Note:
Contact BioLegend Technical Services with any questions regarding cell viability. BioLegend uses an automated cell counter for counting and assessing cell viability, however other methods for assessing cell viability are suitable. For more information about the protocol used by BioLegend please reference “PBMC viability assessment—general methods”.
- Dilute cells in an appropriate volume prior to staining
3.1 Prepare 1 x 106 fresh human PBMCs in 45 μL of Cell Staining Buffer in 12 x 75 mm flow cytometry tubes.
- Block cells
4.1 Add 5 μL of Human TruStain FcX™ Fc blocking reagent to 1 x 106 cells in 45 µL of Cell Staining Buffer (total volume = 50 μL). Incubate for 10 minutes at 4°C.
4.2 While cells are incubating in Fc Block, proceed to step 5, "Stain cells with cell surface antibodies."
- Stain cells with cell surface antibodies
Notes:
If you plan to multiplex samples using TotalSeq™ hashtags, there are two commonly used approaches to staining samples with hashtag antibodies.
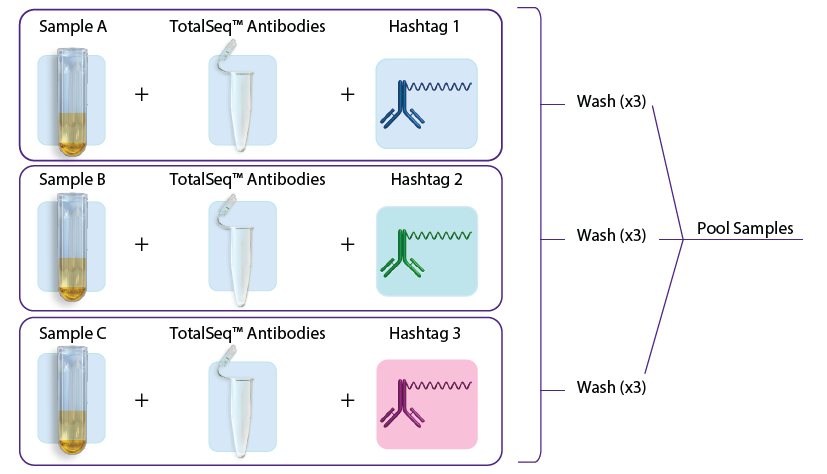
- In the first approach, individual samples are stained with TotalSeq™ antibodies (user created antibody pool) and hashtags in a single step, washed three times to remove unbound antibodies, and subsequently pooled for loading into one or more lanes on the 10x Genomics instrument. See figure 1A.
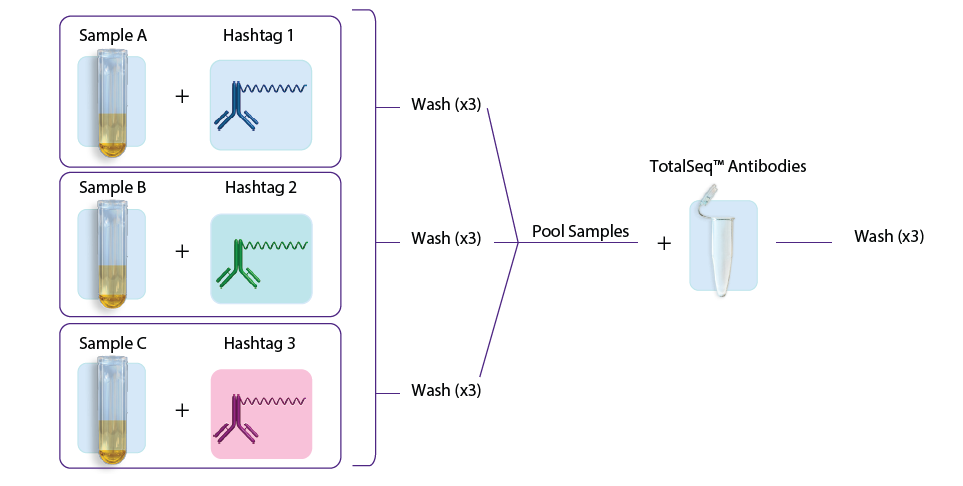
- The second approach involves first staining individual samples with hashtags, washing three times, sample pooling, and then staining with the TotalSeq™ antibodies, followed by another three washes. See figure 1B.
- Whenever possible, we recommend staining TotalSeq™ antibodies and hashtag reagents at the same time, rather than performing sequential staining. Although there are potential cost savings in the second approach because you are staining with a single TotalSeq™ antibody cocktail (such as the Universal Cocktail, or other custom cocktails), the multiple washes can result in increased sample loss and reduced cell viability. Sample washing should not be reduced as it will result in poor quality data. Additional information regarding use of TotalSeq™ hashtag antibodies can be found here: Efficient Multiplexing With TotalSeq™ Hashtags.
A.
B.
Figure 1: A. Staining with TotalSeq™ antibodies and hashtags in a single step before pooling samples. B. Staining individual samples with hashtag reagents followed by pooling and staining with additional TotalSeq™ antibodies in subsequent steps.
5.1 Staining individual samples with TotalSeq™ Hashtag Antibodies for sample pooling
Notes:
- If not using TotalSeq™ Hashtags, or if you are staining cells with a TotalSeq™ antibody pool and hashtags in a single step before pooling samples (figure 1A), proceed to step 5.2.
- If sequentially staining individual samples with hashtag antibodies, pooling samples then staining with a TotalSeq™-B antibody pool, as seen in figure 1B, follow steps below before staining your samples with an antibody pool. Please note if sequentially staining cell recovery may be diminished and cell viability negatively impacted in some cases. If incorporating hashtags into your antibody pool, plan to include an individual hashtag for each sample in step 5.1.1 in the next section.
5.1.1 Make a hashtag staining solution for each sample using titrated amount of each hashtag in Cell Staining Buffer. Add the calculated amount of each hashtag antibody to a low protein binding microcentrifuge tube and bring up to 50 µL total volume with Cell Staining Buffer.
5.1.2 Centrifuge the hashtag solution at 14,000 x g at 2 – 8°C for 10 minutes before adding to the cells to ensure removal of protein aggregates.
5.1.3 Carefully pipette out the prepared hashtag solution, avoiding the bottom of the tube, and add it to the 50 µL blocked cell suspension.
5.1.4 Incubate for 30 minutes at 4°C.
5.1.5 Wash cells, add 3 mL of Cell Staining Buffer, mix by gently pipetting 5 times. Centrifuge at 4°C for 5 minutes at 400 – 600 x g depending on your sample type.
5.1.6 Repeat wash twice for a total of 3 washes.
Note:
It is extremely important to thoroughly decant the wash buffer and resuspend the cell pellet either with pipetting or gentle vortexing. Discard supernatant with a single firm but not overly forceful motion. Proceed to absorb any remaining liquid on the lip of the tube with a clean paper towel.
5.1.7 After the final wash, resuspend each sample in 100 µL of Cell Staining Buffer and verify cell concentration for each sample.
5.1.8 Based on each sample’s cell concentration, combine an equal number of cells from each sample to achieve 1 x 106 cells total in a low protein binding microcentrifuge tube. Mix by gently pipetting.
5.1.9 If volume of combined cells is less than 50 µL, adjust volume with Cell Staining Buffer up to 50 µL to achieve 1 x 106 cells/50 µL. If the final volume of combined cells is greater than 50 µL, centrifuge at 4°C for 5 minutes at 400 – 600 x g. Following centrifugation, adjust volume to 50 µL by removing Cell Staining buffer. After addition or removal of Cell Staining Buffer, gently resuspend cells by pipetting.
Example: If you combined 6 samples at 10 µL of each for a total of 60 µL, after centrifugation you would remove 10 µL for a total volume of 50 µL, resulting in 1 x 106 cells/50 µL.
5.1.10 Proceed with staining your combined samples using section 5.2 "Cell Staining with TotalSeq™ Liquid Antibody Pool and TotalSeq™ Hashtag Antibodies" below. Cells will already be stained with TotalSeq™ Hashtag Antibodies, so do not add additional Hashtag Antibodies to the TotalSeq™-B antigen specific antibody pool.
5.2 Cell Staining with TotalSeq™ Liquid Antibody Pool and TotalSeq™ Hashtag Antibodies
5.2.1 Prepare antibody pool using titrated amounts of each specific TotalSeq™-B antibody. For more information regarding titration of TotalSeq-B™ antibodies please read Tips and Tricks for Titrating TotalSeq™ Antibodies. If you have additional questions, please reach out to BioLegend Tech Services.
5.2.2 When performing dual staining with TotalSeq™-B cell hashtag antibodies and TotalSeq™-B antigen specific antibodies, we recommend adding cell hashing antibodies into each respective sample’s TotalSeq™-B antibody pool (figure 1A).
5.2.3 If the antibody pool volume is less than 50 µL, adjust volume with Cell Staining Buffer up to 50 µL. If the volume of the pool is above 50 µL, no volume adjustment is necessary.
5.2.4 Centrifuge the antibody pool at 14,000 x g at 2 – 8°C for 10 minutes before adding to the cells. This is critical to ensure removal of protein aggregates.
5.2.5 Carefully pipette out the prepared antibody pool, avoiding the bottom of the tube, and add the TotalSeq™ antibody pool to the 50 µL blocked cell suspension.
5.2.6 Incubate for 30 minutes at 4°C.
6. Wash cells
6.1 Add 3 mL of Cell Staining Buffer, mix by gently pipetting 5 times. Centrifuge at 4°C for 5 minutes at 400 – 600 x g depending on your sample type. Repeat step twice for a total of 3 washes.
Note:
It is extremely important to thoroughly decant the wash buffer and resuspend the cell pellet either with pipetting or gentle vortexing. Discard supernatant with a single firm but not overly forceful motion. Proceed to absorb any remaining liquid on the lip of the tube with a clean paper towel.
6.2 After the final wash, decant the supernatant and gently mix the cells by pulse vortexing the cell pellet at medium intensity in the residual volume. Residual volume should be between 50 – 100 µL.
6.3 Proceed immediately to intracellular staining.
Intracellular staining
7. Fixation
7.1 Add 0.5 mL of Fixation Buffer to cells from previous step. Gently mix the cells by pulse vortexing at medium intensity. Fix cells for 20 minutes at room temperature.
7.2 Centrifuge at 1000 x g for 5 minutes, discard supernatant.
8. Permeabilization and blocking
8.1 Dilute Intracellular Staining Perm Wash Buffer (10X) to 1X in deionized water.
8.2 Resuspend fixed cells in 3 mL of Intracellular Staining Perm Wash Buffer, mix by gently pipetting 5 times. Centrifuge at 1000 x g for 5 – 10 minutes. Repeat step twice for a total of 3 washes.
8.3 After the final wash, decant the supernatant and gently mix the cells by pulse vortexing the cell pellet at medium intensity in the residual volume. Residual volume should be between 50 – 100 µL.
8.4 Add 5 µL of True-Stain Monocyte Blocker™, and, 5 µL of Human TruStain FcX™ to the residual volume of cells.
Note:
True-Stain Monocyte Blocker™ is used to reduce non-specific background of intracellular TotalSeq™ conjugates.
8.5 Gently mix the cells by pulse vortexing at medium intensity and incubate for 10 minutes at 4°C.
8.6 During blocking, proceed immediately with steps 9.1 through 9.3.
9. Stain cells with intracellular antibodies
9.1 In a new, clean, low protein binding microcentrifuge tube, prepare the intracellular antibody pool using titrated amounts (up to 1 µg) of each TotalSeq™-B antibody. For more information regarding TotalSeq™ antibody concentrations, please reach out to BioLegend Technical Services.
9.2 After antibodies have been combined, bring the volume up to 100 µL with 1x Intracellular Staining Perm Wash Buffer.
9.3 Centrifuge the antibody pool at 14,000 x g at 2 – 8°C for 10 minutes. This is critical to ensure removal of protein aggregates.
9.4 Following centrifugation, carefully pipette out the antibody pool avoiding the bottom of the tube and add to the cell suspension.
9.5 Stain for 30 minutes at 4°C.
9.6 Add 3 mL of Intracellular Staining Perm Wash Buffer, mix by gently pipetting 5 times. Centrifuge at 4°C for 5 minutes at 1000 x g. Repeat wash 2 more times for a total of 3 washes.
9.7 Resuspend the cells by pulse vortexing the cell pellet at medium intensity in residual volume. Residual volume should be between 50 – 100 µL.
9.8 Slowly filter cells through a 40 µm Flowmi™ cell strainer.
9.9 Verify cell concentration after filtration. Use Buffer EB to make any final adjustments to cell concentration before loading the 10x Chromium.
Note:
Viability indicators will no longer allow discrimination of cells based on membrane integrity, however, Trypan Blue can be used to obtain total cell count.
Proceed to:
Chromium Next GEM Single Cell 3' Reagent Kits v3.1 with Feature Barcoding technology for Cell Surface Protein user guide (CG000317).
Note: Add 2 extra cycles to the 10x cDNA amplification step 2.2.
Recommended Sequencing Depth for Cell Surface Protein Library
To obtain sufficient read coverage for Cell Surface Protein libraries follow recommended library loading and pooling specifications provided in 10x Genomics user guides. See table below for sequencing depth recommendations for Cell Surface Protein libraries, which are applicable to both the Single Cell Gene Expression Solution and the Single Cell Immune Profiling Solution.
| Library Type | Minimum Sequencing Depth |
|---|---|
| Cell Surface and Intracellular Protein Library <100 Antibody Derived Tag (ADT) panel | 5,000 |
| Cell Surface and Intracellular Protein Library ≥100 ADT panel | 10,000 |
Disclaimer: Buyer is solely responsible for determining whether Buyer has all intellectual property rights that are necessary for Buyer's intended uses of the BioLegend TotalSeq™ products. For example, for any technology platform Buyer uses with TotalSeq™, it is Buyer's sole responsibility to determine whether it has all necessary third-party intellectual property rights to use that platform and TotalSeq™ with that platform.
BioLegend is a registered trademark of BioLegend, Inc. All other trademarks are the property of their respective owners.